Labeling carbon backbone amino acid12/24/2023  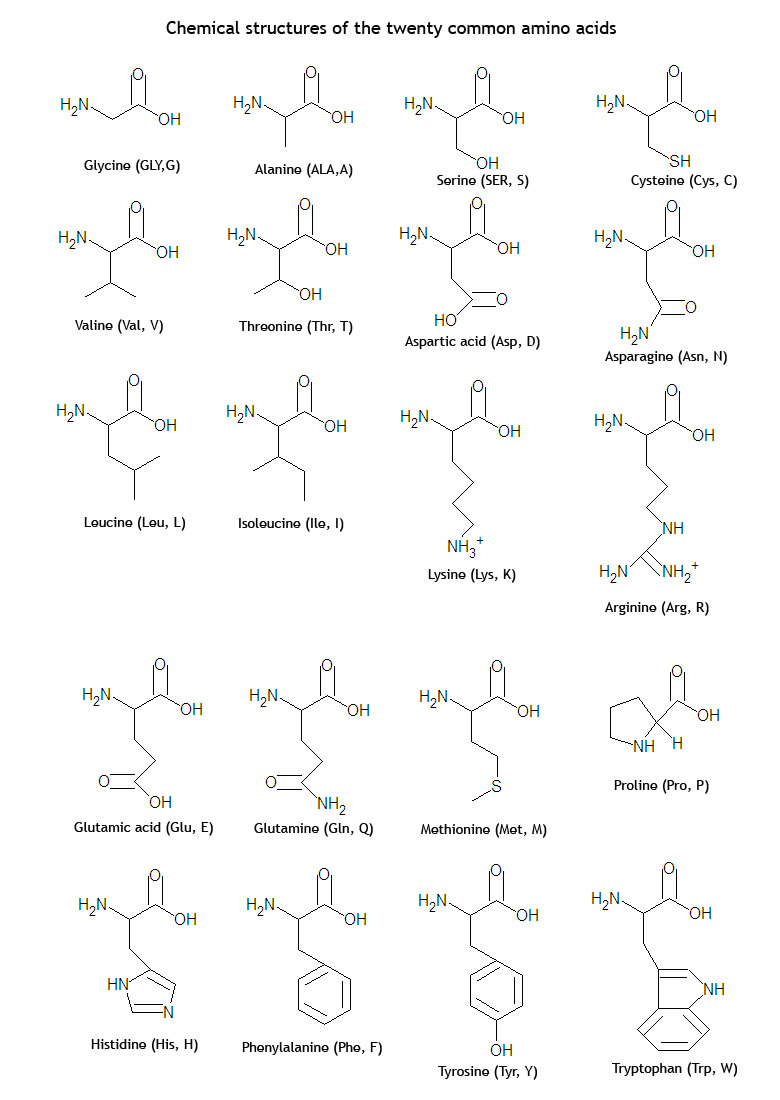
Characterization of the surfactin synthetase C-terminal thioesterase domain as a cyclic depsipeptide synthase. Catalytic role of the condensation domain. Peptide bond formation in nonribosomal peptide biosynthesis. An optimized ATP/PPi-exchange assay in 96-well format for screening of adenylation domains for applications in combinatorial biosynthesis. Mechanisms of β-amino acid incorporation in polyketide macrolactam biosynthesis. The crystal structure of the adenylation enzyme VinN reveals a unique β-amino acid recognition mechanism. Miyanaga, A., Cieślak, J., Shinohara, Y., Kudo, F. Structural basis for the activation of phenylalanine in the non-ribosomal biosynthesis of gramicidin S. A stepwise Huisgen cycloaddition process: copper( I)-catalysed regioselective ‘ligation’ of azides and terminal alkynes. Peptidotriazoles on solid phase: -triazoles by regiospecific copper( I)-catalysed 1,3-dipolar cycloadditions of terminal alkynes to azides. Yeast surface display for screening combinatorial polypeptide libraries. The tyrocidine biosynthesis operon of Bacillus brevis: complete nucleotide sequence and biochemical characterization of functional internal adenylation domains. Folding and function in α/β-peptides: targets and therapeutic applications. Backbone modification of a polypeptide drug alters duration of action in vivo. New open-chain and cyclic tetrapeptides, consisting of α-, β 2-, and β 3-amino-acid residues, as somatostatin mimics-a survey. Biosynthesis of natural products containing β-amino acids. Expanding substrate promiscuity by engineering a novel adenylating-methylating NRPS bifunctional enzyme. Structural elucidation of the bispecificity of A domains as a basis for activating non-natural amino acids. Reprogramming nonribosomal peptide synthetases for ‘clickable’ amino acids. Engineering the substrate specificity of the DhbE adenylation domain by yeast cell surface display. Introduction of a non-natural amino acid into a nonribosomal peptide antibiotic by modification of adenylation domain specificity. Directed evolution of a gatekeeper domain in nonribosomal peptide synthesis. 
Directed evolution of the nonribosomal peptide synthetase AdmK generates new andrimid derivatives in vivo. Computational structure-based redesign of enzyme activity. Exploitation of the selectivity-conferring code of nonribosomal peptide synthetases for the rational design of novel peptide antibiotics. Recent advances in engineering nonribosomal peptide assembly lines. Explorations of catalytic domains in non-ribosomal peptide synthetase enzymology. Assembly-line enzymology for polyketide and nonribosomal peptide antibiotics: logic, machinery, and mechanisms. Molecular mechanisms underlying nonribosomal peptide synthesis: approaches to new antibiotics. Nonribosomal peptide synthesis-principles and prospects. Nonribosomal peptide synthetases involved in the production of medically relevant natural products. Nonribosomal peptides: from genes to products. Reinvigorating natural product combinatorial biosynthesis with synthetic biology. Harnessing the biosynthetic code: combinations, permutations, and mutations. Natural products as sources of new drugs from 1981 to 2014. 
0 Comments
Leave a Reply.AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
